- Facility
Biophysics & Structural Biology (B2S)
- Microcalorimetry, Circular Dichroïsm, Surface Plasmon Resonance, NMR, Chromatography, Light Scattering, Fast kinetics, Cryomicroscopy
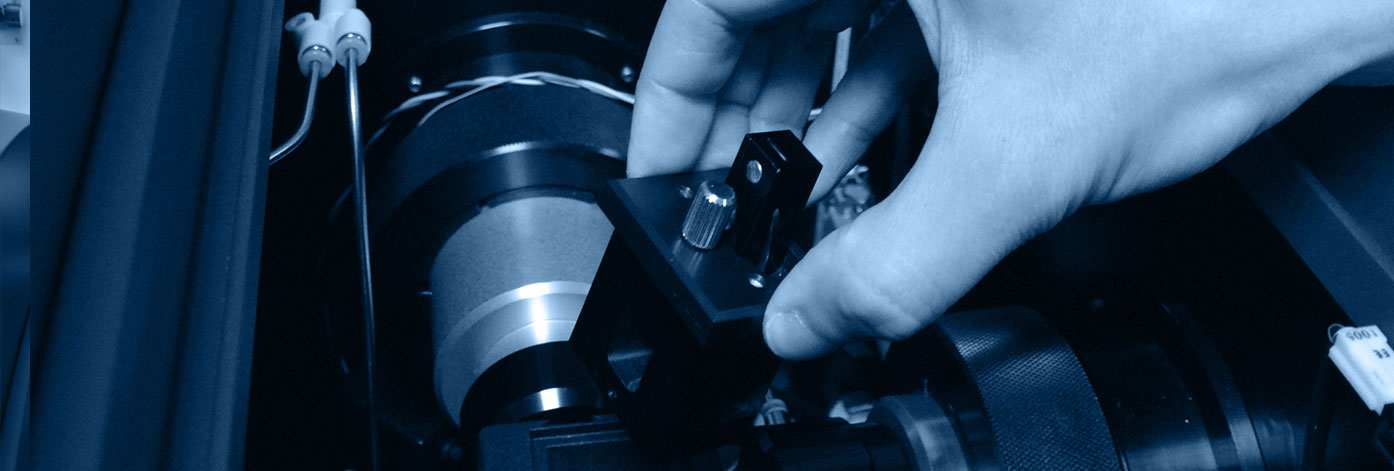
The Biophysics and Structural Biology (B2S) platform offers a combination of equipment for in vitro physicochemical characterization of proteins and interactions. It provides access to several biophysical (circular dichroism, microcalorimetry, surface plasmon resonance, fluorescence) and structural biology (NMR, X-ray) devices, and is based on the skills of the IMoPA research teams. It belongs to the ARBRE association (Association of Resources for Biophysical Research in Europe) created in 2015. It allows the development of multidisciplinary projects to characterize 1) the interactions involved between biological macromolecules and, 2) the 3D structure biological macromolecules.
The B2S staff has a real technical and methodological expertise to address different issues: The stability / characterization of proteins and complexes in solution, the three-dimensional structure of proteins and complexes, fast kinetics, the characterization of protein-protein and protein-ligand interactions as well as the isolation and characterization of a broad spectrum of products by liquid chromatography.
The platform operates in two modes: 1) the service mode (turnkey services along with, or not, collaborations) and 2) the provision of devices for expert users after initial training.
Any new collaborator must contact the person in charge of the Core Facility to define the best strategy to adopt for the project and complete the service request form. Any user will accept the terms and conditions for service use by signing the agreement form.
We are always available for users during the use of the devices and for any advice and assistance.
Each user must retrieve its data at the end of the analyses. In the usual case, the raw acquisition data are kept for 1 year on computer stations. The platform does not guarantee the recovery of data in case of malfunction. The user is responsible for the final data storage.
- Risser F, Collin S, Dos Santos-Morais R, Gruez A, Chagot B, Weissman KJ. Towards improved understanding of intersubunit interactions in modular polyketide biosynthesis: docking in the enacyloxin IIa polyketide synthase. J Struct Biol. 2020 Jul 25.
 10.1016/j.jsb.2020.107581 ,
10.1016/j.jsb.2020.107581 ,  32717326
32717326 Roret T, Alloing G, Girardet JM, Perrot T, Dhalleine T, Couturier J, Frendo P, Didierjean C, Rouhier N. Sinorhizobium meliloti YrbA binds divalent metal cations using two conserved histidines. Biosci Rep. 2020 Oct 30 ; 40(10):BSR20202956.
 10.1042/BSR20202956 ,
10.1042/BSR20202956 ,  32970113
32970113 Rzeigui M, Traikia M, Jouffret L, Kriznik A, Khiari J, Roy O, Taillefumier C. Strengthening Peptoid Helicity through Sequence Site-Specific Positioning of Amide Cis-Inducing NtBu monomers. J Org Chem. 2019 Dec 24.
 10.1021/acs.joc.9b02916 ,
10.1021/acs.joc.9b02916 ,  31873018 ,
31873018 ,  HAL-02430577
HAL-02430577 Ioannou I, Kriznik A, Chekir L, Ghoul M. Effect of the Processing Temperature on the Degradation of Food Flavonoids: Kinetic and Calorimetric Studies on Model Solutions. J. Food Eng. and Technol. 2019 ; 8(2):91-102.
 10.32732/jfet.2019.8.2.91 ,
10.32732/jfet.2019.8.2.91 ,  HAL-02392716
HAL-02392716 Chagot ME, Dos Santos Morais R, Dermouche S, Lefebvre D, Manival X, Chipot C, Dehez F, Quinternet M. Binding properties of the quaternary assembly protein SPAG1. Biochem J. 2019 May 22.
 10.1042/BCJ20190198 ,
10.1042/BCJ20190198 ,  31118266 ,
31118266 ,  HAL-02140524
HAL-02140524 S Ahmed Zennia S, Mati A, Charron C, Cakir-Kiefer C, Kriznik A, Girardet JM. Effect of nonenzymatic deamidation on the structure stability of Camelus dromedarius alpha-lactalbumin.Food Chemistry. 2019 Avril 8.
 10.1016/j.foodchem.2019.04.033 ,
10.1016/j.foodchem.2019.04.033 ,  HAL-02095313
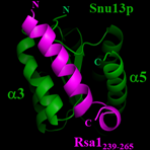
HAL-02095313 Chagot ME, Quinternet M, Rothé B, Charpentier B, Coutant J, Manival X, Lebars I. The yeast C/D box snoRNA U14 adopts a "weak" K-turn like conformation recognized by the Snu13 core protein in solution. Biochimie. 2019 Mar 23.
 10.1016/j.biochi.2019.03.014 ,
10.1016/j.biochi.2019.03.014 ,  30914254 ,
30914254 ,  HAL-02082477
HAL-02082477 Rahuel-Clermont S, Bchini R, Barbe S, Boutserin S, André I, Talfournier F. Enzyme Active Site Loop Revealed as a Gatekeeper for Cofactor Flip by Targeted Molecular Dynamics Simulations and FRET-Based Kinetics. ACS Catal. 2019 ; 9 ; 1337−1346.
 10.1021/acscatal.8b03951 ,
10.1021/acscatal.8b03951 ,  HAL-02022828
HAL-02022828 Yakavets I, Millard M, Lamy L, Francois A, Scheglmann D, Wiehe A, Lassalle H-P, Zorin V, Bezdetnaya L. Matryoshka-Type Liposomes Offer the Improved Delivery of Temoporfin to Tumor Spheroids. Cancers. 2019 September 13.
 10.3390/cancers11091366
10.3390/cancers11091366 de Guillen K, Lorrain C, Tsan P, Barthe P, Petre B, Saveleva N, Rouhier N, Duplessis S, Padilla A, Hecker A. Structural genomics applied to the rust fungus Melampsora larici-populina reveals two candidate effector proteins adopting cystine knot and NTF2-like protein folds. Sci Rep. 2019;9(1):18084.
 10.1038/s41598-019-53816-9 ,
10.1038/s41598-019-53816-9 ,  PMC6889267
PMC6889267