- Facility
Epitranscriptomics & Sequencing (EpiRNA-Seq)
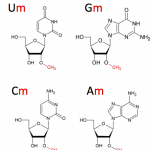
- Next-Generation Sequencing, RNA, RNA modifications

The Epitranscriptomics and Sequencing (EpiRNA-Seq) Core Facility is built on the expertise of the UMR7365 IMoPA in the field of RNA molecular biology to promote its main collaborative services.
Located on the Brabois-Santé campus of the University of Lorraine in Nancy, it provides to the academic and industrial scientific community high-tech resources (Illumina next generation sequencers). It is part of the European COST EPITRAN network and regularly organises training activities.
The core facility has recognized technical and methodological know-how to contribute to various projects in Epitranscriptomics (RiboMethSeq, AlkAniline-Seq, HydraPsiSeq) to address various scientific questions such as dynamics of post-transcriptional modifications in RNAs.
The EpiRNA-Seq Core Facility operates mainly in collaborative mode (involvement of members of the core facility in the steps of your sequencing project from the conception of the experimental design to the statistical analysis of the data) but the delivery mode (mainly sequencing of libraries prepared by you without involvement of the platform) is also possible under certain conditions. The EpiRNA-Seq Core Facility has a differential pricing system (Université de Lorraine members, academics and industrialists).
The EpiRNA-Seq Core Facility is open to the academic and industrial scientific community at the local, national and international level.
The team is at your disposal to help you by bringing its technical and methodological know-how and to follow you from A to Z in your sequencing projects.
The EpiRNA-Seq Core Facility has a differential pricing system (Université de Lorraine members, academic and industrial).
Any new collaborator must first contact the head of the core facility using the "Request form" to define the best strategy to adopt for conducting his study, and obtain a quote. Any user will agree to accept the "terms and conditions". The guidelines to prepare and send samples to the Core Facility are given in "Sample sheet".
Henry BA, Marchand V, Schlegel BT, Helm M, Motorin Y, Lee N. Pseudouridylation of Epstein-Barr Virus Noncoding RNA EBER2 Facilitates Lytic Replication. RNA. 2022 Sep 13:rna.079219.122
 10.1261/rna.079219.122 ,
10.1261/rna.079219.122 ,  36100352 ,
36100352 ,  HAL-03778341
HAL-03778341 Hertler J, Slama K, Schober B, Özrendeci Z, Marchand V, Motorin Y, Helm M. Synthesis of point-modified mRNA. Nucleic Acids Res. 2022 Sep 5:gkac719.
 10.1093/nar/gkac719 ,
10.1093/nar/gkac719 ,  36062567 ,
36062567 ,  HAL-03778358
HAL-03778358 - Dreggors-Walker RE, Cohen LN, Khoshnevis S, Marchand V, Motorin Y, Ghalei H. Studies of mutations of assembly factor Hit 1 in budding yeast suggest translation defects as the molecular basis for PEHO syndrome. J Biol Chem. 2022 Jul 14:102261.
 10.1016/j.jbc.2022.102261 ,
10.1016/j.jbc.2022.102261 ,  35843310 ,
35843310 ,  HAL-03727002
HAL-03727002 - Brégeon D, Pecqueur L, Toubdji S, Sudol C, Lombard M, Fontecave M, de Crécy-Lagard V, Motorin Y, Helm M, Hamdane D. Dihydrouridine in the Transcriptome: New Life for This Ancient RNA Chemical Modification. ACS Chem Biol. 2022 Jun 23.
 10.1021/acschembio.2c00307 ,
10.1021/acschembio.2c00307 ,  35737906 ,
35737906 ,  HAL-03708292
HAL-03708292 Helm M, Motorin Y. Phosphorylation found inside RNA. Nature. 2022 May;605(7909):234-235.
 10.1038/d41586-022-01021-6 ,
10.1038/d41586-022-01021-6 ,  35478019 ,
35478019 ,  HAL-03710903
HAL-03710903 - Pichot F, Marchand V, Helm M, Motorin Y. Machine learning algorithm for precise prediction of 2'-O-methylation (Nm) sites from experimental RiboMethSeq datasets. Methods. 2022 Mar 18:S1046-2023(22)00074-3.
 10.1016/j.ymeth.2022.03.007 ,
10.1016/j.ymeth.2022.03.007 ,  35314341 ,
35314341 ,  HAL-03616347
HAL-03616347 Khoshnevis S, Dreggors-Walker RE, Marchand V, Motorin Y, Ghalei H. Ribosomal RNA 2'-O-methylations regulate translation by impacting ribosome dynamics. Proc Natl Acad Sci U S A. 2022 Mar 22 ; 119(12):e2117334119.
 10.1073/pnas.2117334119 ,
10.1073/pnas.2117334119 ,  35294285 ,
35294285 ,  HAL-03612834
HAL-03612834 - Delhermite J, Tafforeau L, Sharma S, Marchand V, Wacheul L, Lattuca R, Desiderio S, Motorin Y, Bellefroid E, Lafontaine DLJ. Systematic mapping of rRNA 2'-O methylation during frog development and involvement of the methyltransferase Fibrillarin in eye and craniofacial development in Xenopus laevis. PLoS Genet. 2022 Jan 18;18(1):e1010012.
 10.1371/journal.pgen.1010012 ,
10.1371/journal.pgen.1010012 ,  35041640 ,
35041640 ,  HAL-03537766
HAL-03537766 - Ragon M, Bertheau L, Dumont J, Bellanger T, Grosselin M, Basu M, Pourcelot E, Horrigue W, Denimal E, Marin A, Vaucher B, Berland A, Lepoivre C, Dupont S, Beney L, Davey H, Guyot S. The Yin-Yang of the Green Fluorescent Protein: Impact on Saccharomyces cerevisiae stress resistance. J Photochem Photobiol B. 2022 Nov 24;238:112603.
 10.1016/j.jphotobiol.2022.112603 ,
10.1016/j.jphotobiol.2022.112603 ,  36459911
36459911 Schöller E, Marks J, Marchand V, Bruckmann A, Powell CA, Reichold M, Mutti CD, Dettmer K, Feederle R, Hüttelmaier S, Helm M, Oefner P, Minczuk M, Motorin Y, Hafner M, Meister G. Balancing of mitochondrial translation through METTL8-mediated m3C modification of mitochondrial tRNAs. Mol Cell. 2021 Nov 8:S1097-2765(21)00910-2.
 10.1016/j.molcel.2021.10.018 ,
10.1016/j.molcel.2021.10.018 ,  34774131 ,
34774131 ,  HAL-03429971
HAL-03429971

Formation CNRS Entreprises
( -> )
The EpiRNA-Seq Core Facility organizes the following training school (CNRS Formation Entreprises) : Epitranscriptomics: mapping and analysis of RNA modifications by Next-Generation Sequencing. This training school will be held from 17st to 19st May 2021 on the Core Facility in Nancy. For more details : https://cnrsformation.cnrs.fr/epitranscriptomics-mapping-and-analysis-of-rna-modifications-by-next-generation-sequencing?axe=140

Epitranscriptomics training school (COST Epitran)
( -> )
The NGS Core Facility in collaboration with Pr Iouri Motorine (UMR7365 IMoPA) is organizing a training school on Epitranscriptomics with the support of COST Epitran from 11-14 february 2019 in Nancy (France). This training is dedicated to young phD students or postdocs. The session in 2019 is closed. The next session will take place in February 2020, please send an email to the head of EpiRNA-Seq core facility. For more information : /sites/default/files/users/Projets/Diapositive1.png